Research on differential expression of hepatic stress protein after reduced-size liver transplantation in rats
-
摘要:
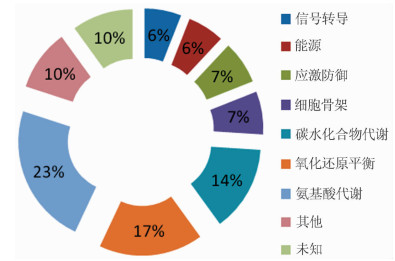
目的 探讨大鼠减体积肝移植术后肝脏应激蛋白的差异表达情况。 方法 分别将改良大鼠减体积肝移植模型术后1、3、7 d的肝脏组织标本的双向电泳图谱与供体和受体原肝脏组织的双向电泳图谱进行比较, 以变化倍数大于10倍或小于1/10为标准选择差异蛋白点, 再利用质谱技术及数据库对差异表达的蛋白质点进行分析和鉴定。 结果 共发现了72个差异表达的蛋白质点, 通过质谱分析和一系列的鉴定, 最终发现有32种功能明确的蛋白, 其中热休克蛋白-8与肥厚激动剂反应蛋白在减体积移植术后肝脏表达差异较大, 占所有差异蛋白质点的7%(5/72)。 结论 在大鼠减体积肝移植术后移植肝脏中发现参与肝脏应激反应的蛋白质, 为下一步研究这些蛋白与肝移植术后肝脏缺血-再灌注损伤的关系提供了前期研究数据。 Abstract:Objective To discuss the differential expression of hepatic stress proteins after reduced-size liver transplantation in rats. Methods The specimens of liver tissues were procured on 1 d, 3 d and 7 d after the improved model of reduced-size liver transplantation in rats. Then, the two-dimensional electrophoresis of these specimens was compared with that of the original liver tissues of normal donors and recipients. The differentially expressed protein spots were selected with the standard of change times greater than 10 or less than 1/10 and then were analyzed and identified by mass-spectrometric technique and data bases. Results Seventy-two differentially expressed protein spots were found in total. And the 32 kinds of proteins were identified with definite function through mass spectrometry and a series of identifications. The expression difference of heat shock protein-8 and hypertrophy agonist reactive protein was larger, amounting 7% (5/72) of all differential proteins. Conclusions This study provides fundamental research data for studying the relation between liver ischemia-reperfusion injury after liver transplant and the above differential proteins of stress reaction in transplant liver which are found after reduced-size liver transplantation in rats. -
Key words:
- Reduced-size liver transplantation /
- Rat /
- Stress reaction /
- Mass spectrometry
-
表 1 大鼠减体积肝移植术后移植肝脏中发现的差异表达的应激蛋白
Table 1. Differential expression of stress proteins found in transplant liver after reduce-size liver transplantation in rats
自编号 蛋白名称 基因 功能分类 Accession No. 蛋白质PI 1 热休克蛋白-8 HSP-8 应激反应蛋白 gi|13242237 5.37 2 肥厚激动剂反应蛋白 IAH1 应激反应蛋白 gi|34786935 6.33 注:Accession No.为基因库GenBank的登录号;PI为等电点(isoelectric point) -
[1] Brewis IA, Brennan P. Proteomics technologies for the global identification and quantification of proteins[J]. Adv Protein Chem Struct Biol, 2010, 80:1-44. doi: 10.1016/B978-0-12-381264-3.00001-1 [2] Müller SA, Findeiβ S, Pernitzsch SR, et al. Identification of new protein coding sequences and signal peptidase cleavage sites of Helicobacter pylori strain26695 by proteogenomics[J]. J Proteomics, 2013, 86:27-42. doi: 10.1016/j.jprot.2013.04.036 [3] Armengaud J, Hartmann EM, Bland C. Proteogenomics for environmental microbiology[J]. Proteomics, 2013, 13(18/19):2731-2742. http://www.ncbi.nlm.nih.gov/pubmed/23636904 [4] Armengaud J, Trapp J, Pible O, et al. Non-model organisms, a species endangered by proteogenomics[J]. J Proteomics, 2014, 105:5-18. doi: 10.1016/j.jprot.2014.01.007 [5] Sun H, Chen C, Shi M, et al. Integration of mass spectrometry and RNA-Seq data to confirm human ab initio predicted genes and lncRNAs[J]. Proteomics, 2014, 14(23/24):2760-2768. https://www.researchgate.net/profile/Lu_Xie4/publication/267275973_Integration_of_mass_spectrometry_and_RNA-Seq_data_to_confirm_human_ab_initio_predicted_genes_and_lncRNAs/links/54c6edd10cf22d626a35e77f.pdf?origin=publication_detail [6] Gustafsson OJ, Arentz G, Hoffmann P. Proteomic developments in the analysis of formalin-fixed tissue[J]. Biochim Biophys Acta, 2014, DOI: 10.1016/j.bbapap.2014.10.003[Epub ahead of print]. [7] Vincenti DC, Murray GI. The proteomics of formalin-fixed wax-embedded tissue[J]. Clin Biochem, 2013, 46(6):546-551. doi: 10.1016/j.clinbiochem.2012.10.002 [8] Kim SC, Page EK, Knechtle SJ. Urine proteomics in kidney transplantation[J]. Transplant Rev, 2014, 28(1):15-20. doi: 10.1016/j.trre.2013.10.004 [9] Sigdel TK, Salomonis N, Nicora CD, et al. The identification of novel potential injury mechanisms and candidate biomarkers in renal allograft rejection by quantitative proteomics[J]. Mol Cell Proteomics, 2014, 13(2):621-631. doi: 10.1074/mcp.M113.030577 [10] Kornasiewicz O, Bojarczuk K, Bugajski M, et al. Application of a proteomic approach to identify proteins associated with primary graft non-function after liver transplantation[J]. Int J Mol Med, 2012, 30(4):755-764. https://www.spandidos-publications.com/ijmm/30/4/755/abstract [11] 刘静, 李江, 张升宁, 等.改良法构建大鼠减体积肝移植模型的建立[J].中国组织工程研究与临床康复杂志, 2010, 14(18):3252-3257. http://mall.cnki.net/magazine/article/xdkf201018012.htmLiu J, Li J, Zhang SN, et al. Modified model of reduced-size liver transplantation in rats[J]. J Clin Rehabil Tissue Eng Res, 2010, 14(18):3252-3257. http://mall.cnki.net/magazine/article/xdkf201018012.htm [12] Amacher DE. The discovery and development of proteomic safety biomarkers for the detection of drug-induced liver toxicity[J]. Toxicol Appl Pharmacol, 2010, 245(1):134-142. doi: 10.1016/j.taap.2010.02.011 [13] Franchin C, Cesaro L, Pinna LA, et al. Identification of the PLK2-dependent phosphopeptidome by quantitative proteomics[J]. PLoS One, 2014, 9(10):e111018. doi: 10.1371/journal.pone.0111018 [14] Kesharwani SS, Nandekar PP, Pragyan P, et al. Comparative proteomics among cytochrome p450 family 1 for differential substrate specificity[J]. Protein J, 2014, 33(6):536-548. doi: 10.1007/s10930-014-9586-6 [15] Wei J, Zhang F, Zhang Y, et al. Proteomic investigation of signatures for geniposide-induced hepatotoxicity[J]. J Proteome Res, 2014, 13(12):5724-5733. doi: 10.1021/pr5007119 [16] Bassols A, Costa C, Eckersall PD, et al. The pig as an animal model for human pathologies:a proteomics perspective[J]. Proteomics Clin Appl, 2014, 8(9/10):715-731. https://www.researchgate.net/publication/264500034_The_pig_as_an_animal_model_for_human_pathologies_A_proteomics_perspective [17] Bohra R, Klepacki J, Klawitter J, et al. Proteomics and metabolomics in renal transplantation-quo vadis?[J]. Transpl Int, 2013, 26(3):225-241. doi: 10.1111/tri.2013.26.issue-3 [18] Akbarian A, Michiels J, Golian A, et al. Gene expression of heat shock protein 70 and antioxidant enzymes, oxidative status, and meat oxidative stability of cyclically heat-challenged finishing broilers fed Origanum compactum and Curcuma xanthorrhiza essential oils[J]. Poult Sci, 2014, 93(8):1930-1941. doi: 10.3382/ps.2014-03896 [19] Yamamoto TM, Wang L, Fisher LA, et al. Regulation of Greatwall kinase by protein stabilization and nuclear localization[J].Cell Cycle, 2014, 13(22):3565-3575. doi: 10.4161/15384101.2014.962942 [20] Leung AM, Redlak MJ, Miller TA. Role of heat shock proteins in oxygen radical-induced gastric apoptosis[J].J Surg Res, 2014, DOI:10.1016/j.jss. 2014.07.013[Epub ahead of print]. [21] Zhu J, Zhou Y, Wang GN, et al. Cell cycle arrest, apoptosis and autophagy induced by iminosugars on K562 cells[J]. Eur J Pharmacol, 2014, 731:65-72. doi: 10.1016/j.ejphar.2014.03.013 [22] Wang R, Shao F, Liu Z, et al. The Hsp90 inhibitor SNX-2112, induces apoptosis in multidrug resistant K562/ADR cells through suppression of Akt/NF-κB and disruption of mitochondria-dependent pathways[J]. Chem Biol Interact, 2013, 205(1):1-10. doi: 10.1016/j.cbi.2013.06.007 [23] Steel R, Cross RS, Ellis SL, et al. Hsp70 architecture:the formation of novel polymeric structures of Hsp70.1 and Hsc70 after proteotoxic stress[J]. PLoS One, 2012, 7(12):e52351. doi: 10.1371/journal.pone.0052351 [24] Kim JY, Yenari MA, Lee JE. Regulation of inflammatory transcription factors by heat shock protein 70 in primary cultured astrocytes exposed to oxygen-glucose deprivation[J]. Neuroscience, 2014, DOI: 10.1016/j.neuroscience.2014.11.057[Epub ahead of print]. [25] Silva PN, Furuya TK, Braga IL, et al. Analysis of HSPA8 and HSPA9 mRNA expression and promoter methylation in the brain and blood of Alzheimer's disease patients[J]. J Alzheimers Dis, 2014, 38(1):165-170. https://www.ncbi.nlm.nih.gov/pubmed/23948933/ [26] Boswell-Casteel RC, Johnson JM, Duggan KD, et al. Overproduction and biophysical characterization of human HSP70 proteins[J]. Protein Expr Purif, 2014, DOI: 10.1016/j.pep.2014.09.013[Epub ahead of print]. -




 下载:
下载: